-Search query
-Search result
Showing all 43 items for (author: harvey & sc)

EMDB-36241: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (consensus map)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-36242: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (Masked refinement of the cap domain)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-36243: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (masked refinement of the transmembrane domain)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-36244: 
Cryo-EM structure of mouse Piezo1-MDFIC(C240A) complex
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-35577: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (composite map)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

PDB-8imz: 
Cryo-EM structure of mouse Piezo1-MDFIC complex (composite map)
Method: single particle / : Zhou Z, Ma X, Lin Y, Cheng D, Bavi N, Li JV, Sutton D, Yao M, Harvey N, Corry B, Zhang Y, Cox CD

EMDB-28092: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-093
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28090: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-040
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO

EMDB-28091: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-045
Method: single particle / : Li H, Callaway H, Yu X, Shek J, Saphire EO
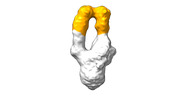
EMDB-28093: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-156
Method: single particle / : Shek J, Callaway H, Li H, Yu X, Saphire EO
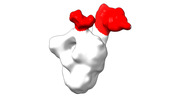
EMDB-28094: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-234
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28095: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-260
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28096: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-279
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28097: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-290
Method: single particle / : Yu X, Callaway H, Li H, Shek J, Saphire EO
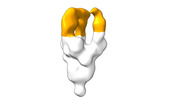
EMDB-28098: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-294
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28099: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-295
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28100: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-299
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28102: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-334
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
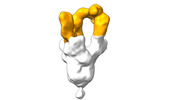
EMDB-28103: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-360
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28104: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-361
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28105: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-362
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28106: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-368
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28168: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-292
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28169: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-333
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO
EMDB-28170: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-355
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-28171: 
Negative stain EM map of SARS-CoV-2 Spike in complex with CoVIC-371
Method: single particle / : Callaway H, Li H, Yu X, Shek J, Saphire EO

EMDB-11997: 
Structure of a nanoparticle for a COVID-19 vaccine candidate
Method: single particle / : Duyvesteyn HME, Stuart DI

PDB-7b3y: 
Structure of a nanoparticle for a COVID-19 vaccine candidate
Method: single particle / : Duyvesteyn HME, Stuart DI

EMDB-22143: 
Cryo-EM structure of NusG-CTD bound to 70S ribosome
Method: single particle / : Washburn R, Zuber P

PDB-6xe0: 
Cryo-EM structure of NusG-CTD bound to 70S ribosome (30S: NusG-CTD fragment)
Method: single particle / : Washburn R, Zuber P, Sun M, Hashem Y, Shen B, Li W, Harvey S, Acosta-Reyes FJ, Knauer SH, Frank J, Gottesman ME

EMDB-20224: 
BG505 SOSIP.664 with 2G12 Fab2
Method: single particle / : Cottrell CA, de Val N, Ward AB

PDB-6ozc: 
BG505 SOSIP.664 with 2G12 Fab2
Method: single particle / : Cottrell CA, Ward AB

PDB-4v47: 
Real space refined coordinates of the 30S and 50S subunits fitted into the low resolution cryo-EM map of the EF-G.GTP state of E. coli 70S ribosome
Method: single particle / : Gao H, Sengupta J, Valle M, Korostelev A, Eswar N, Stagg SM, Van Roey P, Agrawal RK, Harvey ST, Sali A, Chapman MS, Frank J

PDB-4v48: 
Real space refined coordinates of the 30S and 50S subunits fitted into the low resolution cryo-EM map of the initiation-like state of E. coli 70S ribosome
Method: single particle / : Gao H, Sengupta J, Valle M, Korostelev A, Eswar N, Stagg SM, Van Roey P, Agrawal RK, Harvey ST, Sali A, Chapman MS, Frank J

EMDB-1395: 
Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron microscopy.
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

EMDB-1055: 
Locking and unlocking of ribosomal motions.
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Hervey SC, Ehrenberg M, Frank J

EMDB-1056: 
Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron microscopy.
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

PDB-1qza: 
Coordinates of the A/T site tRNA model fitted into the cryo-EM map of EF-Tu ternary complex (GDP.Kirromycin) bound 70S ribosome
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

PDB-1qzb: 
Coordinates of the A-site tRNA model fitted into the cryo-EM map of 70S ribosome in the pre-translocational state
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

PDB-1qzc: 
Coordinates of S12, SH44, LH69 and SRL separately fitted into the cryo-EM map of EF-Tu ternary complex (GDP.Kirromycin) bound 70S ribosome
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

PDB-1qzd: 
EF-Tu.kirromycin coordinates fitted into the cryo-EM map of EF-Tu ternary complex (GDP.Kirromycin) bound 70S ribosome
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

PDB-1r2w: 
Coordinates of L11 with 58nts of 23S rRNA fitted into the cryo-EM map of the 70S ribosome
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J

PDB-1r2x: 
Coordinates of L11 with 58nts of 23S rRNA fitted into the cryo-EM map of EF-Tu ternary complex (GDP.Kirromycin) bound 70S ribosome
Method: single particle / : Valle M, Zavialov A, Li W, Stagg SM, Sengupta J, Nielsen RC, Nissen P, Harvey SC, Ehrenberg M, Frank J
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
